Specializing in Single Cell and Spatial Multiomics
AccuSomatic Amplification for Single Cell Sequencing enables the discovery of true somatic single nucleotide variations (SNVs) in single cells that could not be accurately detected by any prior method. The existing methods suffer from multiple sources of artifacts resulting in more than 20,000 false somatic SNV calls per cell and >90% false-positive results.
AccuSomatic Amplification eliminates >99% of errors in somatic SNV calls while maintaining the same detection sensitivity. Our unique service enables researchers to unlock the full potential of single-cell sequencing in disease research and drug development for the first time.
Single-cell RNA-Seq enables the study of cell-to-cell transcriptome heterogeneity, which opens a new field in biology. Singulomics Corporation is highly experienced in a high-throughput single cell and single nucleus RNA sequencing and data analysis using the 10x Genomics Chromium platform for fresh/frozen cells and frozen tissue samples.
Our workflow incorporates dead cell removal or nuclei isolation depending on the sample type and quality to ensure optimal results. We have successfully conducted single nucleus RNA-Seq for a wide variety of tissue types and tissue biopsies. To enable the discovery of single-cell transcriptomes at a deeper level, we also offer deep single-cell RNA-Seq services for individual cells and ultra-low-input RNA samples.
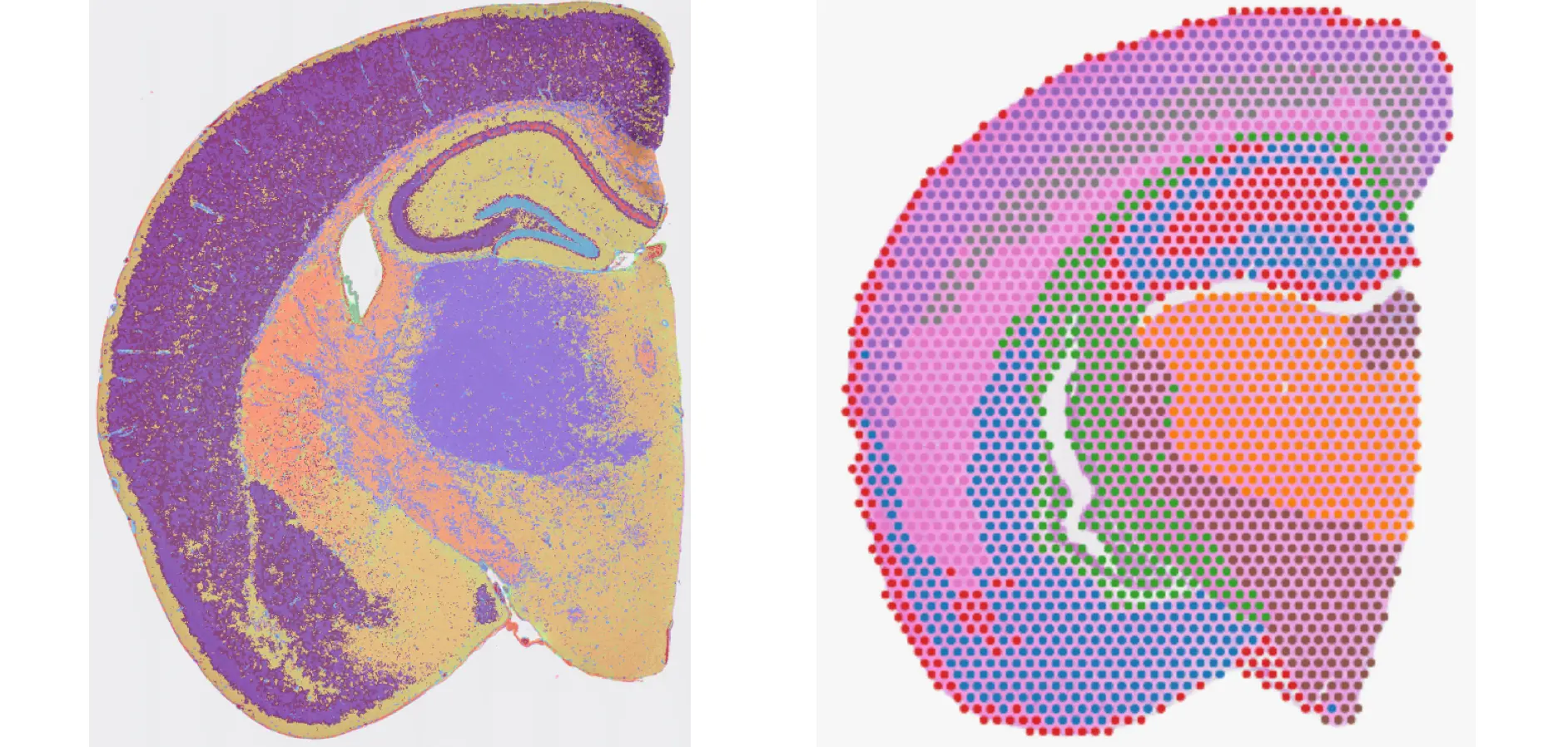
At SingulOmics, we offer spatial transcriptomic analysis using 10x Genomics’ Visium and Visium HD Spatial Gene Expression workflows to map the whole transcriptome across FFPE and frozen tissue sections with morphological context and high cellular resolution—achieving single-cell scale resolution with Visium HD. We also support spatial multiomic analysis using the Immunofluorescence workflow for simultaneous protein detection. Our expert team, with deep experience in histology, spatial and single-cell technologies, and bioinformatics analysis, delivers comprehensive end-to-end services—from tissue sectioning to publication-ready analysis results.
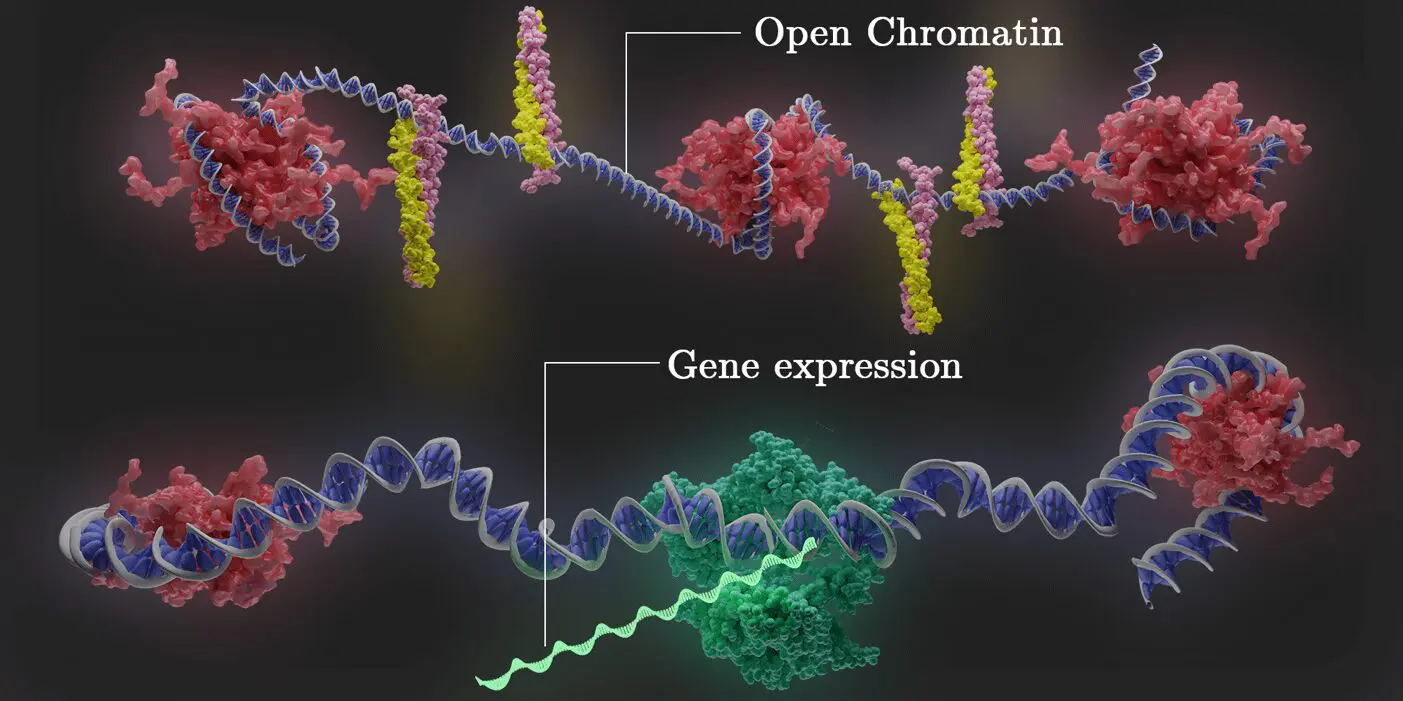
Discover how genes are expressed and regulated using our 10x Single Cell Multiome ATAC + Gene Expression service. With simultaneous profiling of gene expression and open chromatin from the same cell, you can link input regulatory signals with their output gene expression at the single-cell level for thousands of cells. We leverage years of experience in nuclei isolation from cryopreserved cells and flash-frozen tissues to provide accurate, high-quality data.
Single-cell ATAC Sequencing is a technique used to determine genome-wide chromatin accessibility at the single-cell level. With single-cell ATAC-Seq, chromatin profiling of large numbers of nuclei can be performed in parallel, resulting in fast, accurate epigenomic profiling. Using the industry-leading Chromium Single-Cell ATAC Solution from 10x Genomics, our PhD-level scientists and bioinformaticians can analyze thousands of nuclei per run at single-cell resolution. In combination with Single-Cell RNA-Seq, our end-to-end services and integrated analysis enable researchers to link input regulatory signals with their output gene expression at the single-cell level.
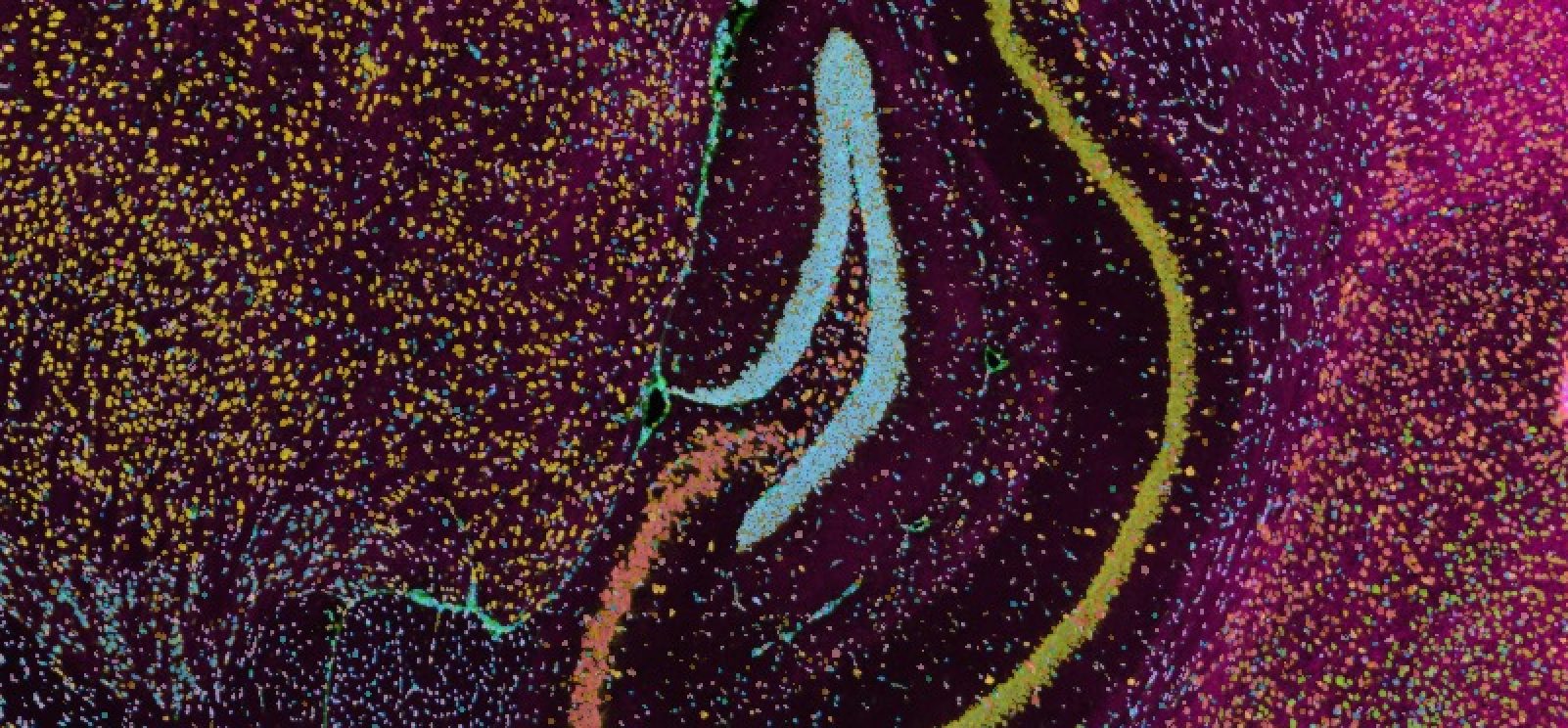
The Xenium platform provides high-plex, in situ RNA detection with subcellular resolution, generating precise spatial gene expression maps from FFPE or fresh-frozen tissues. Singulomics offers a streamlined Xenium workflow—from tissue sectioning to high-resolution imaging—delivering detailed spatial transcript information directly from your samples. Xenium panels range from focused targets of a few hundred genes to the Xenium 5K panel with over 5,000 genes, with options for custom additions. Combined with Visium HD or single-cell RNA-seq, our Xenium service enables multi-scale insights that link global tissue architecture with subcellular transcript localization.